Students Name: Paul Ephraim D, Thirupuasundari E, Janardhanan D
Nucleus segmentation is a critical task in biomedical image analysis, underpinning applications from disease diagnosis to cellular research. Although conventional convolutional neural networks (CNNs) have delivered notable success in this field, their reliance on local receptive fields limits their ability to capture long-range dependencies and complex spatial relationships. In response, we propose an innovative segmentation framework that integrates a self-attention mechanism within a hybrid architecture, termed Swin-UNet. Our model incorporates Swin Transformer blocks into the encoder, leveraging a hierarchical self-attention mechanism to dynamically model both local fine-grained features and global contextual relationships. This self-attention module enables the network to efficiently aggregate information across disparate regions of the image, overcoming the inherent limitations of traditional CNNs in handling spatial intricacies. The effective combination of self-attention and convolutional operations not only enhances feature extraction but also maintains computational efficiency by processing high-resolution inputs with linear complexity.
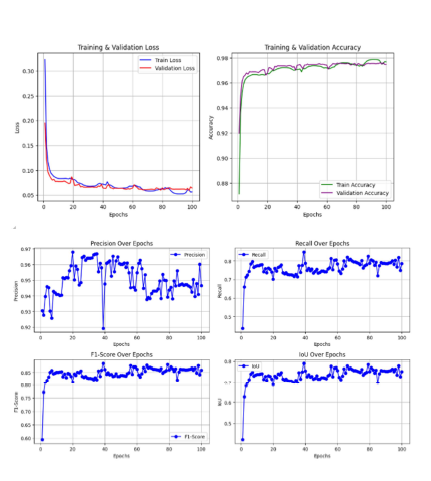
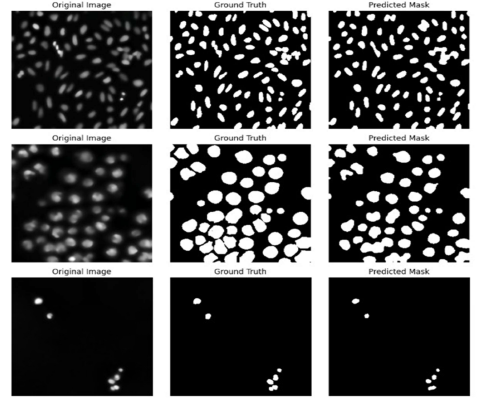
Trained on a diverse set of microscopy images, the proposed architecture demonstrates robustness under varied imaging conditions and improves segmentation accuracy. Extensive evaluations reveal that Swin-UNet, with its integrated self-attention strategy, outperforms standard CNN-based approaches, making it a promising solution for nucleus segmentation and a wide array of biomedical image analysis tasks.
